OUR RESEARCH
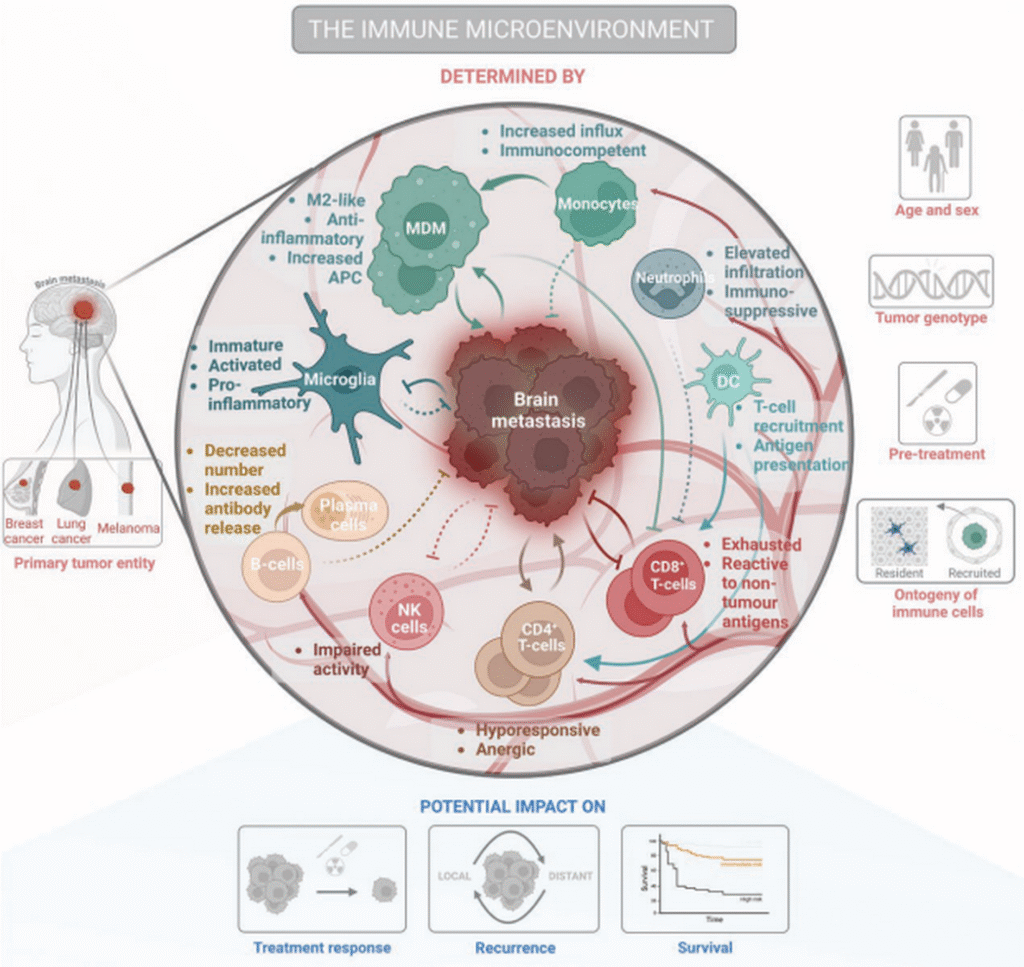
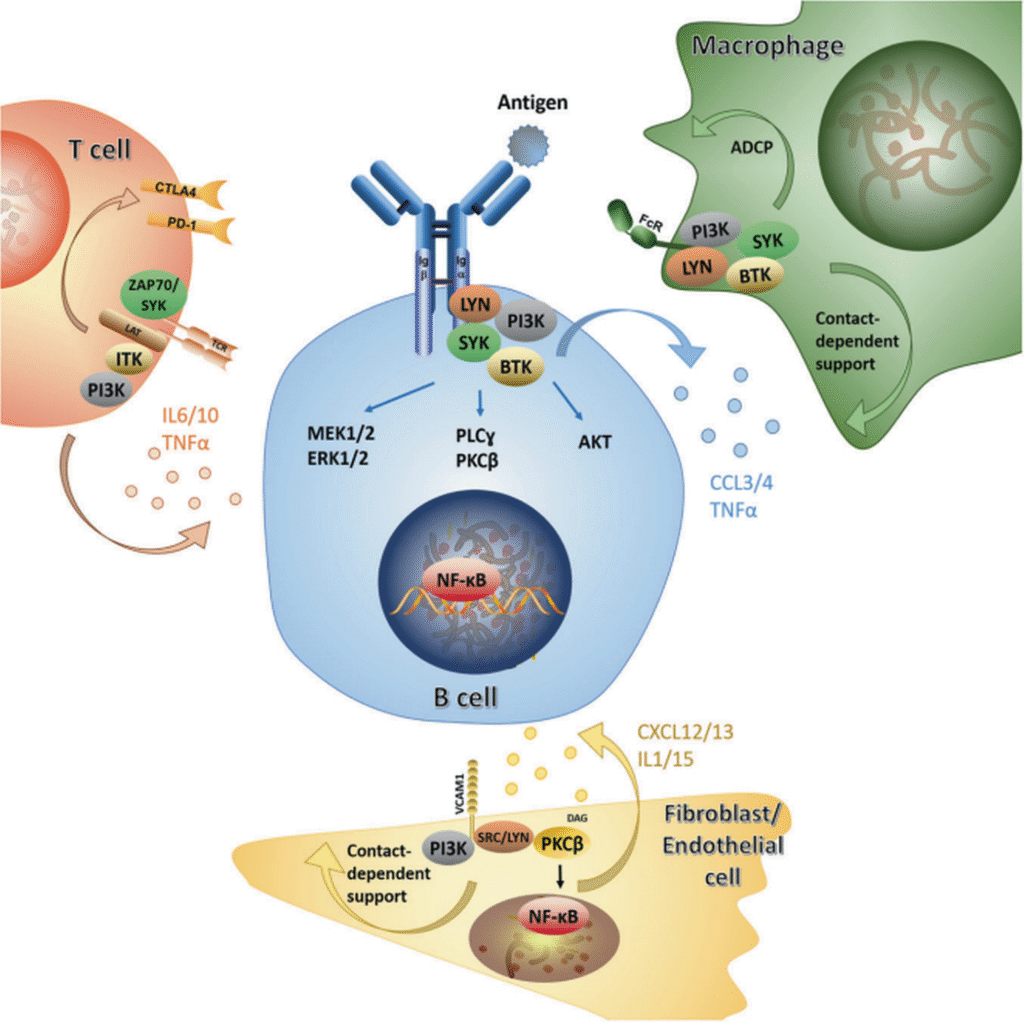
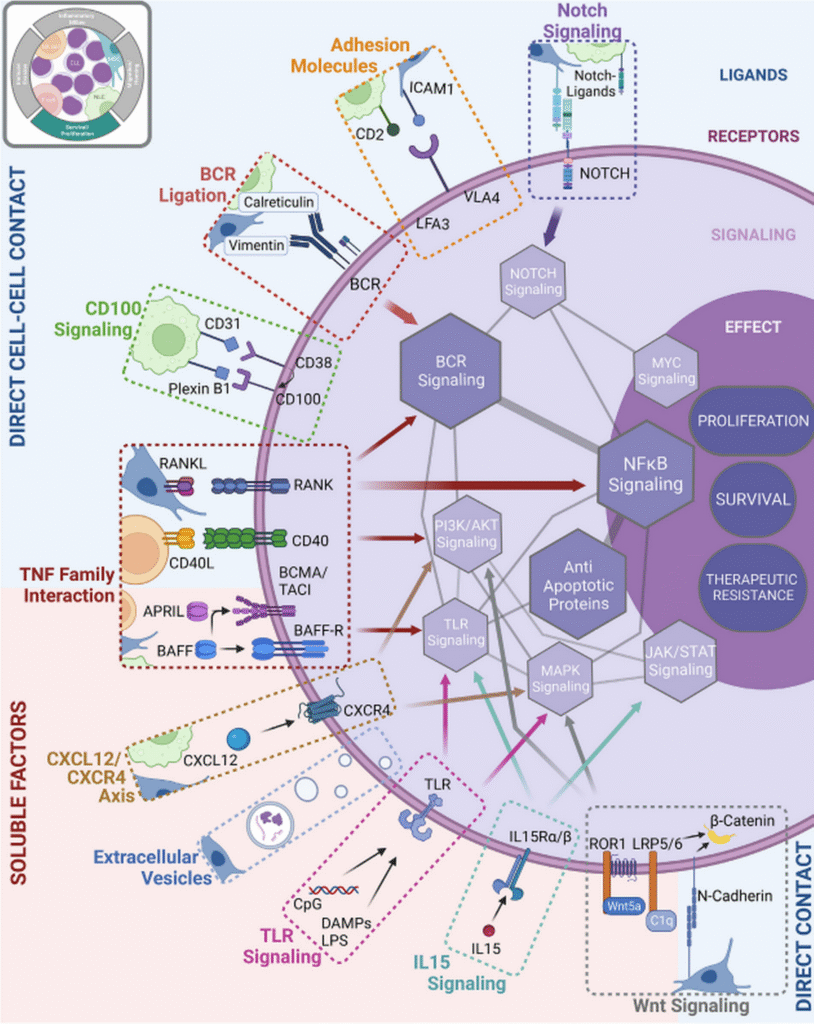
The conceptual framework of cancer hallmarks has expanded considerably in recent decades. Cancer is no longer perceived merely as a disease of malignant cells, but as a complex tissue composed of tumor cells intimately interacting with their surrounding tumor microenvironment (TME). Within this complex ecosystem, multiple cell types engage in dynamic, multidirectional interactions that collectively drive malignant progression and influence therapeutic outcomes.
Our primary research goal is to elucidate the reciprocal molecular and cellular interactions occurring between tumor cells and their host environment, particularly the immune cells. To achieve this, we employ
- Advanced and integrative methodologies, including single-cell analytics to unravel cellular heterogeneity and signaling complexity
- Functional high-content knockout screens for systematic target identification and validation
- 3D multiculture models that accurately recapitulate tumor architecture and cell-cell interactions
- Genetically engineered mouse models that authentically represent cancer progression and immune interactions in vivo
Through this comprehensive approach, we aim to uncover fundamental principles underlying tumor-host interdependence. By dissecting the precise signaling pathways, transcriptional responses, and cellular dynamics governing these interactions, our group seeks to profoundly advance our mechanistic understanding of cancer biology. Ultimately, this will inform the development of innovative and personalized therapeutic combinations, significantly improving clinical outcomes for patients.